AAVカプシド指向性進化
組み換えアデノ随伴ウイルス(AAV)は、その広範な組織指向性、導入遺伝子の長期間の発現能、非病原性、および低い免疫原性によって遺伝子治療およびワクチン用途の遺伝子デリバリーベクターとして重宝されています。
しかし、遺伝子治療ベクター開発が盛んになるにつれ、既存のAAVセロタイプ(血清型)では、遺伝子治療ツールとしてのポテンシャルがある程度制限されているのが現状です。第一に、既存のAAVセロタイプは様々な組織をターゲットできるとなっていますが、往々にして臨床応用では、既存のセロタイプでは特異的にターゲットできない組織や細胞を治療対象していることがあります。 第二に、治療対象組織に対して、1種または複数のセロタイプでターゲットできる場合もありますが、遺伝子デリバリー効率が低すぎる場合があること、また複数の組織に対して指向性をもつため、標的以外の組織にもオフターゲット効果をもたらし、特異性に劣ることもあります。 第三に、すでに特定のAAVセロタイプに対する中和抗体を持つ患者がいるため、中和抗体にブロックされAAVが使用できない可能性があることや、初回のAAVの投与後、患者が中和抗体を持ってしまい、2回目以降のAAV投与が不可能になる場合があること。最後に、一部のセロタイプは、高タイター、純度、および安定的に製造することが本質的に困難であること、などです。 これらの制限を克服するために、AAVカプシドに自然界での進化を模倣させて改変し、新しいAAVバリアントの開発を実現させるのがこの受託サービスの特長です。
指向性進化は、生体分子の機能増強に広く用いられているハイスループットアプローチで、進化遺伝学的な多様化と選択による自然淘汰のプロセスを模倣しています。 AAVキャプシドの指向性進化は、野生型AAVキャプシッド遺伝子を変異させ、非常に多様なAAVキャプシドライブラリーを構築することから始めます。次に、これらのライブラリーをスクリーニングして、目的の機能が増強された新規キャプシド変異型を同定します。指向性進化はキャプシッドタンパク質の構造と機能の関係についての事前知識を必要としないため、機能から合理的に設計するキャプシッドエンジニアリングに比べて研究者に好まれています。
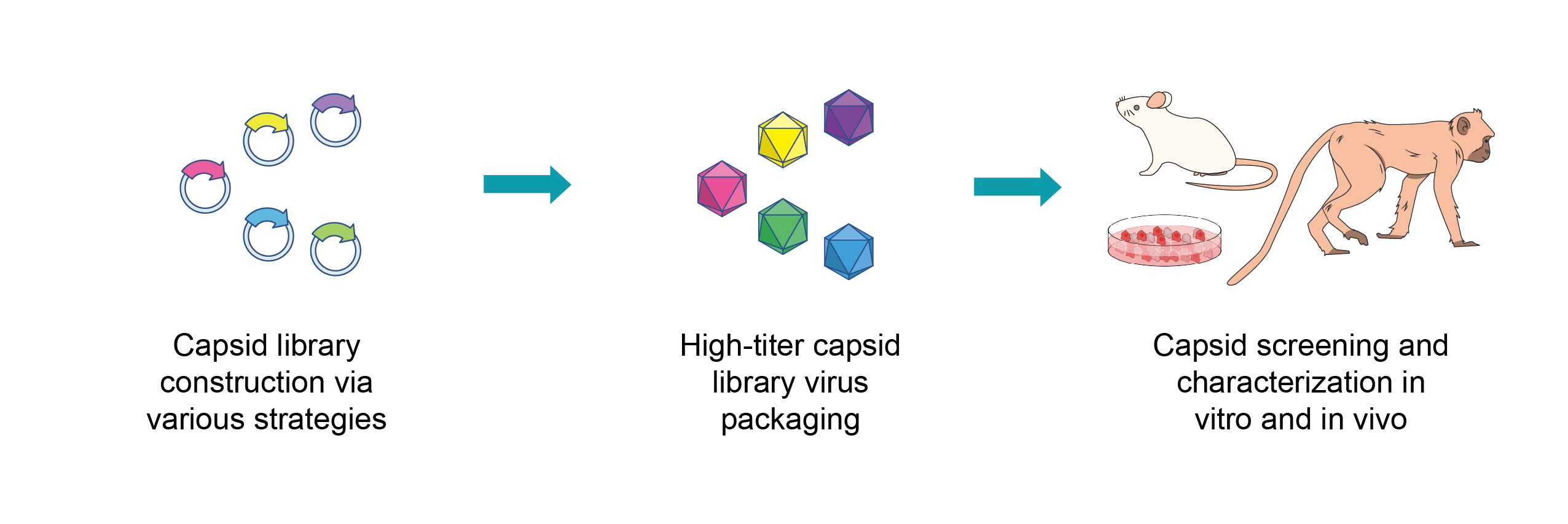
このサービスの特長
- フルサービスプラットフォーム: VectorBuilderは、基礎研究など前臨床研究用AAV作製、臨床用AAVのCRO(受託企業)そしてCDMO(受託開発製造機関)として、研究ベンチから患者のベッドサイドまで必要なAAVの作成サービスをワンストップで提供できる世界で唯一の受託サービス企業です。当社のAAV受託サービスは、 ベクターデザインと最適化, ベクタークローニング, ライブラリー構築, ウイルスパッケージング, キャプシッドの指向性進化と合理的エンジニアリング, AAV生体局在のプロファイリング, そして GMPグレード製造を提供しています。
- 多様なアプローチによる高複雑なキャプシッドライブラリーの構築: 当社はすでにプール型ライブラリー構築に関して豊富な知識と経験を持っています。そのため、変異導入方法や複数ベクター構築方法を組み合わせた非常に複雑なキャプシッドライブラリーのデザインを行い、ライブラリー構築します。
- 高タイターのキャプシッドライブラリー用ウイルスパッケージング: キャプシドライブラリーは、1ステップまたは2ステップのウイルスパッケージングによって、各ウイルス粒子がウイルスゲノムにキャプシド変異型を含むようにし、また高いウイルスタイターで納品できるように構築します。
- 非ヒト霊長類(NHP)を含む複数の動物種でのIn vivoスクリーニング: 当社は、in vitroスクリーニングの豊富な実績に加えて、マウス、ラット、そしてカニクイザル(Macaca fascicularis;別名cynomolgus monkey)とアカゲザル (Macaca Mulatta)の2種の非ヒト霊長類をin vivoスクリーニングプラットフォームで提供しています。 In vivoスクリーニングは高度な訓練を受けた専属技術員らによってAAALAC認定施設にて実施しています。
- 完全な技術サポート: 経験豊富な当社の研究者が、ライブラリー構築からin vivoスクリーニング、ライブラリの設計からNGS分析まで、AAVキャプシドプロジェクトのあらゆる側面をカバーする包括的な技術サポートを提供します。
カプシド指向性進化とスクリーニングのワークフロー
AAVキャプシド進化のワークフロー全体における最も重要なステップは、rep遺伝子とキャプシド遺伝子変異型からなるキメラAAVゲノムを運ぶプラスミドDNAからなる非常に多様なAAVキャプシドライブラリーの構築です。 キャプシド遺伝子変異型は、エラープローンPCR、ランダムペプチドディスプレイ、DNAファミリーシャッフリング、またはin silicoデザインなど、さまざまなアプローチを使用して効率的に構築できます。 次に、キャプシドライブラリーはウイルス粒子にパッケージングされ、各ウイルス粒子はキメラゲノムキャプシド変異体を保持します。次にウイルスライブラリーは、キメラAAVの特性をテストするスクリーニングを行います。 1) 特定の組織や器官内の目的細胞に対して効率的に遺伝子導入を行えるかどうか、または 2) 細胞特異的な受容体に高アフィニティーで結合できるか、または 3) 中和抗体の回避能があるかどうか。スクリーニングに合格したウイルスゲノムは、標的細胞から回収され、2回目のスクリーニング用の小ライブラリーに集められます。信頼性の高いヒットを単離するするために、通常、数ラウンドのスクリーニングが実行されます。 次に、得られたヒットを検証し特性の評価を行い、目的の特性が強化された新規AAVキャプシド変異型として確立します(図 1)。
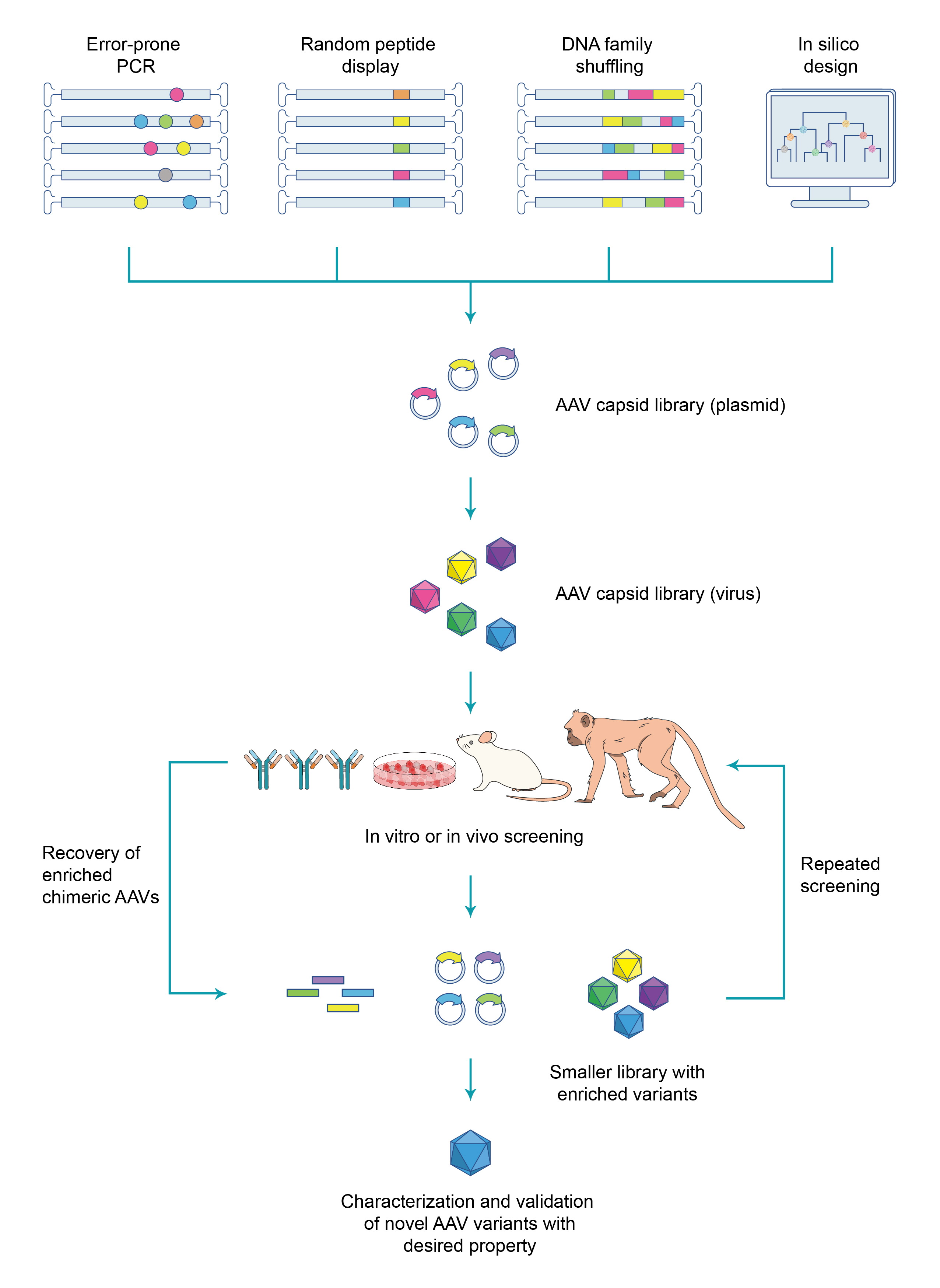
図 1. 指向性進化(directed evolution)による新規AAVカプシドスクリーニングの一般的なワークフロー
カプシドライブラリー構築
以下のストラテジーは、さまざまなAAVキャプシド変異型を作成するために一般的に使用されている方法です。個々の キャプシド変異型をAAVベクターに大規模に並行してクローニングし、キメラAAVゲノム(子のゲノムは各プラスミドベクターが正常型rep遺伝子と変異型キャプシド遺伝子を1対1で持つ)を形成し、スクリーニング用のキャプシドライブラリーを構築します。
エラープローン PCR
Error-prone PCRは、標準的なPCR法を使って、AAVキャプシド遺伝子にランダムに変異誘発を行います。この方法は、非常多くの様々なAAVキャプシドライブラリーを開発するための最も簡単なアプローチです。エラープローンPCRは、精度の低いポリメラーゼ、より長い伸長時間、より高いMg2 +濃度、Mn2 +の添加、ランダムな点突然変異を導入するためにさまざまな濃度でdNTP濃度を添加して、最適化の劣るPCR条件を組み合わせて実行します。
ランダムペプチドディスプレイ
ランダムペプチドディスプレイでは、ウイルスの自然な細胞相互作用を変化させ、特定の細胞受容体に再標的化することを目的として、通常7〜9アミノ酸からなるランダムペプチド配列をAAVキャプシドの特定の部位に挿入します。 ペプチドは通常、ペプチドの表面露出を促進し、ウイルスと宿主の相互作用にも重要なAAVキャプシドの位置に挿入されます。たとえば、AAV2キャプシドの587位と588位(可変領域VRIII内)は、ほとんどのAAV2ベースのペプチドディスプレイライブラリに適した挿入部位です。その領域にペプチドを挿入すると、AAV2のヘパラン硫酸プロテオグリカン(HSPG、プライマリAAV2受容体)モチーフが無効になるためです。そして提示されたペプチドが細胞表面分子と効率的に相互作用することを可能にします。
DNAファミリーシャッフリング
DNAファミリーシャッフリングは、異なるAAVセロタイプに由来する親キャプシド遺伝子を分子異種交配によってキメラAAVキャプシドを精製する非常に効率的なアプローチです。さまざまなAAVセロタイプの親キャプシド遺伝子を断片化し、部分的な配列相同性に基づいてそれらを再結合するプライマーを必要としないPCRによって、新規の完全長キャプシド変異型を再構築します。またこの代替方法として、合理的設計(AAVの生物的知識に基づいてキャプシドを変更する)と指向性進化を組み合わせた合成シャッフリングによって、複雑度の高いライブラリーを作成することもできます。このアプローチでは、突然変異誘発に適したキャプシドの位置が、自然に存在するAAVセロタイプの詳細な構造および配列分析に基づいて特定、検証されます。 変異を含むフラグメントを合成し、完全長の新規キャプシド変異体を形成するためにアッセンブルされます。
イン シリコデザイン
In silico AAVキャプシドライブラリーデザインは、キャプシド変異体配列の計算予測にさまざまなアプローチを利用しており、AAVパフォーマンスの向上に貢献します。一般的に使用されるアプローチの1つは、推定上の祖先AAVライブラリーのin silico設計と、それに続く、親和性が改善された非常に強力な祖先キャプシド配列を特定するための実験的検証を含む祖先再構築です。 このアプローチの背後にある理論的根拠は、自然淘汰のプロセスを生き延びて出現した進化的AAV中間体は、ウイルスの構造と機能を維持しながら、独自の特性を持っている可能性が高いということです。マシーンラーニングは、仮想のキャプシド変異体から実行可能なウイルス産生の可能性を予測するために計算アルゴリズム使用する、もう1つのよく使用されるin silicoデザイン方法です。マシーンラーニングのアルゴリズムは、利用可能な入力データに大きく依存してタンパク質の構造と機能の関係を学習し、それを適用してウイルスキャプシドアセンブリなどの複雑な生理学的プロセスの結果を予測します。
カプシドライブラリーのウイルスパッケージング
キャプシドライブラリのウイルスパッケージングは、1ステップまたは2ステップのプロセスで作製されます:
カプシドライブラリーのワンステップ(one-step)パッケージング
AAVキャプシドライブラリーをパッケージングするための従来のアプローチは、パッケージング細胞がキャプシドライブラリーおよびアデノウイルスヘルパープラスミドで共トランスフェクトされるワンステップ方法です。 この方法は広く使われていますが、クロスパッケージング(キャプシド変異ゲノムとキャプシドが一致しないAAV粒子の生成)とキャプシドモザイク現象(異なるゲノムに由来するキャプシドタンパク質から生じるモザイクキャプシドを持つAAV粒子の生成)の欠点があります。これらを克服するために、パッケージング細胞を非常に低いプラスミドライブラリー対細胞比でトランスフェクトして、細胞あたり最大1つの単一ライブラリープラスミドが取り込まれるようにします。
カプシドライブラリーのツーステップ(two-step)パッケージング
ツーステップのパッケージング方法は、最初に野生型キャプシド遺伝子をコードしているがウイルスITRを欠いているヘルパープラスミドとともにパッケージング細胞にコトランスフェクトしてキャプシッドライブラリーを構築します。 この結果、部分的に野生型キャプシドで構成された、AAVトランスファーシャトルと呼ばれるモザイクキャプシドをもつAAV粒子が生成されます。 次に、AAVトランスファーシャトルを低MOIでパッケージング細胞に導入して、細胞あたり最大1つのウイルス粒子が感染するようにします。続いてパッケージング細胞をアデノウイルスで重感染させ、最終的に高力価のウイルスキャプシドライブラリーを生成します。
In vitroとin vivo カプシドスクリーニング
ウイルスキャプシドライブラリーは通常、スクリーニングの目的に基づいて、in vitroまたはin vivoで数ラウンドのスクリーニングが実施され、望ましい特性を備えたキメラAAVを選択します:
In vitro 選択
確立された細胞株をAAVキャプシドライブラリーの選択に利用することは、特に受容体ターゲティング能力が変化したAAV変異型を同定する際によく使用されている方法です。 AAVライブラリーのin vitro選択は迅速かつ技術的に簡単ですが、いくつかの課題があります。 第一に、in vitroで高い形質導入効率に最適化されたベクターは、in vivoで使用した場合に同じ効率を再現できない可能性があります。 第二に、in vitroで高度な標的細胞特異性を示すAAVベクターは、in vivoで翻訳されたときに非標的組織に形質導入する可能性があります。AAVライブラリーをin vitroで選択する他の方法は、特に免疫回避特性を持つ変異型を同定するために、ライブラリーを標的細胞に加える前に免疫化動物からの強力な血清にさらすことです。 ただし、AAVバリアントの免疫応答は、さまざまな要因によりin vivoで翻訳された場合に変化する可能性があります(たとえば、同じAAVベクターの免疫認識は、異なる経路でデリバリーされた場合に変化する可能性があります)。
In vivo選択
In vivo動物モデルは、AAVライブラリーをスクリーニングするためのより信頼性の高いプラットフォームです。特に、培養では増殖できない繊細な細胞型を形質導入できるAAV変異型、または複雑な組織構造を持つ特定の細胞型を形質導入できるAAV変異型を特定します。 In vivoでの選択は、AAVバリアントに関連する潜在的なオフターゲット効果を特定するのにも役立ちます。 マウスとNHPの両方がAAVライブラリーのin vivo選択に広く使用されていますが、NHPモデルは、ヒトとの類似性が高いため、改良されたAAVベクターをスクリーニングするための最も臨床的に適切なプラットフォームです。




